Testing Sensor based behavior: Difference between revisions
No edit summary |
No edit summary |
||
| (13 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
= Sensor based behavior information for functional traits with focus on rumination = | = Sensor based behavior information for functional traits with focus on rumination = | ||
== Part 1 | == Part 1: General introduction == | ||
=== Background and aim of the guideline === | === Background and aim of the guideline === | ||
Recent advancements in sensor technologies have significantly enhanced the capacity to technically support farmers and their advisors in monitoring the health, performance, and welfare of dairy cattle. As presented in the systematic review by Stygar et al. (2021)<ref name="Stygar2021">Stygar, A.H., Gómez, Y., Berteselli, G.V., Dalla Costa, E., Canali, E., Niemi, J.K., Llonch, P., Pastell, M. 2021. A systematic review on commercially available and validated sensor technologies for welfare assessment of dairy cattle. Frontiers in Veterinary Science 8, 177</ref> and in other focused reviews (e.g., Hogeveen et al., 2021), a wide range of commercially available sensor systems exists and promises significant gains in the understanding and improvement of welfare in livestock. The technologies cover the spectrum from wearable devices with multiple functions (e.g., tracking of physiological parameters) to environmental sensors that monitor housing and climatic conditions, and collectively aim to provide actionable insights about animal health, reproductive status and welfare. Most wearable sensors rely on 3D accelerometers, which measure acceleration or motion to quantify cow behaviour. Sensor technology providers use algorithms and pattern recognition to enhance the raw accelerometer data and produce sensor systems which recognize rumination, eating, lying, standing, and other behaviours, using the data from sensors on the cow’s leg, neck, ear, or tail. The integration of sensor systems in livestock farming presents numerous opportunities to enhance animal health, performance and welfare, supporting farmer decision making on individual cow and group level and farm efficiency. However, while large amounts of sensor data are being collected, only a small fraction is currently used on farms, in genetic evaluation and breeding programs, or along the dairy value chain (Brito et al., 2025). To increase confidence in the use of data from advanced technologies and sensor-based herd management systems among key stakeholders (farmers and consultants, authorities, dairy processors, breeding and genetics organizations, and consumers), sensor-derived data need to be combined with routinely recorded data. At present, only a low fraction of commercially available sensor systems is independently validated for welfare assessment (14%; Stygar et al., 2021)<ref name="Stygar2021"/>, and beyond farmers’ own experience, few studies investigated the performance of some sensor systems in diverse farming environments, across different farm and management systems and geographical locations. These challenges motivate the need for coordinated guidance on how to define, process, and use sensor-derived behavioural information. | Recent advancements in sensor technologies have significantly enhanced the capacity to technically support farmers and their advisors in monitoring the health, performance, and welfare of dairy cattle. As presented in the systematic review by Stygar et al. (2021)<ref name="Stygar2021">Stygar, A.H., Gómez, Y., Berteselli, G.V., Dalla Costa, E., Canali, E., Niemi, J.K., Llonch, P., Pastell, M. 2021. A systematic review on commercially available and validated sensor technologies for welfare assessment of dairy cattle. Frontiers in Veterinary Science 8, 177</ref> and in other focused reviews (e.g., Hogeveen et al., 2021), a wide range of commercially available sensor systems exists and promises significant gains in the understanding and improvement of welfare in livestock. The technologies cover the spectrum from wearable devices with multiple functions (e.g., tracking of physiological parameters) to environmental sensors that monitor housing and climatic conditions, and collectively aim to provide actionable insights about animal health, reproductive status and welfare. Most wearable sensors rely on 3D accelerometers, which measure acceleration or motion to quantify cow behaviour. Sensor technology providers use algorithms and pattern recognition to enhance the raw accelerometer data and produce sensor systems which recognize rumination, eating, lying, standing, and other behaviours, using the data from sensors on the cow’s leg, neck, ear, or tail. The integration of sensor systems in livestock farming presents numerous opportunities to enhance animal health, performance and welfare, supporting farmer decision making on individual cow and group level and farm efficiency. However, while large amounts of sensor data are being collected, only a small fraction is currently used on farms, in genetic evaluation and breeding programs, or along the dairy value chain (Brito et al., 2025)<ref name="Brito2025">Poppe, M., H.A. Mulder, M.L. van Pelt, E. Mullaart, H. Hogeveen, and R.F. Veerkamp. 2022. Development of resilience indicator traits based on daily step count data for dairy cattle breeding. Genetics Selection Evolution 54:21. doi:10.1186/s12711-022-00713-x</ref>. To increase confidence in the use of data from advanced technologies and sensor-based herd management systems among key stakeholders (farmers and consultants, authorities, dairy processors, breeding and genetics organizations, and consumers), sensor-derived data need to be combined with routinely recorded data. At present, only a low fraction of commercially available sensor systems is independently validated for welfare assessment (14%; Stygar et al., 2021)<ref name="Stygar2021"/>, and beyond farmers’ own experience, few studies investigated the performance of some sensor systems in diverse farming environments, across different farm and management systems and geographical locations. These challenges motivate the need for coordinated guidance on how to define, process, and use sensor-derived behavioural information. | ||
Against this background, the International Committee of Animal Recording (ICAR) and the International Dairy Federation (IDF) started a joint initiative aiming at improved usability of data across sensor systems and applications. The initiative leaders are the ICAR Functional Traits Working Group (ICAR FTWG) and the IDF Standing Committee of Animal Health and Welfare (IDF SCAHW) in collaboration with international experts from academia and industry organizations. The primary aim of this initiative is to promote the integrated use of sensor data and derived novel traits along the dairy value chain. Standardisation and harmonisation will be supported through guidelines that include basic definitions and recommendations regarding data processing and use. Priorities of work are based on results from a survey with manufacturers and feedback on stakeholder needs. These are: | Against this background, the International Committee of Animal Recording (ICAR) and the International Dairy Federation (IDF) started a joint initiative aiming at improved usability of data across sensor systems and applications. The initiative leaders are the ICAR Functional Traits Working Group (ICAR FTWG) and the IDF Standing Committee of Animal Health and Welfare (IDF SCAHW) in collaboration with international experts from academia and industry organizations. The primary aim of this initiative is to promote the integrated use of sensor data and derived novel traits along the dairy value chain. Standardisation and harmonisation will be supported through guidelines that include basic definitions and recommendations regarding data processing and use. Priorities of work are based on results from a survey with manufacturers and feedback on stakeholder needs. These are: | ||
| Line 18: | Line 18: | ||
The current guideline has the focus on data from sensor systems measuring animal behaviours. These sensor systems can provide information on behavioural measurements like rumination, eating, lying or indexes like activity indexes or alarms like for calving, in heat or health events. Various sensor systems are based on different technologies using different algorithms and provide different information to the farmer. | The current guideline has the focus on data from sensor systems measuring animal behaviours. These sensor systems can provide information on behavioural measurements like rumination, eating, lying or indexes like activity indexes or alarms like for calving, in heat or health events. Various sensor systems are based on different technologies using different algorithms and provide different information to the farmer. | ||
= Part 2: Terminology = | == Part 2: Terminology == | ||
=== Suggested Key Performance Indicators (KPIs) for sensor-based rumination data === | === Suggested Key Performance Indicators (KPIs) for sensor-based rumination data === | ||
| Line 44: | Line 44: | ||
Welch, J. G. 1982. Rumination, particle size and passage from the rumen. J. Anim. Sci. 54:885–894. https:// doi .org/ 10 .2527/ jas1982.544885x. | Welch, J. G. 1982. Rumination, particle size and passage from the rumen. J. Anim. Sci. 54:885–894. https:// doi .org/ 10 .2527/ jas1982.544885x. | ||
= Part 3: Sensor data cleaning = | == Part 3: Sensor data cleaning == | ||
=== Recommendations for data cleaning === | === Recommendations for data cleaning === | ||
| Line 70: | Line 70: | ||
The items in this summary checklist correspond to and summarise the five-step framework described below and are intended as a quick user guide to the more detailed explanations. | The items in this summary checklist correspond to and summarise the five-step framework described below and are intended as a quick user guide to the more detailed explanations. | ||
=== Five-step framework for cleaning sensor data including | === Five-step framework for cleaning sensor data including === | ||
These instructions are proposed by Schodl et al. 2024<ref name="Schodl2024">Schodl, K., Stygar, A., Steininger, F., & Egger-Danner, C., 2024a. Sensor data cleaning for applications in dairy herd management and breeding. Front. Anim. Sci., 5, p.1444948. <nowiki>https://doi.org/10.3389/fanim.2024.1444948</nowiki>.</ref>.) | |||
# '''Verification of the data''' preprocessing: Accurate alignment between animal identifiers and sensor data is critical. Errors such as duplicate device assignments to one animal (or vice versa), broken sensors, and time zone mismatches must be identified and corrected, if possible. It is recommended to consult with digital technology companies for information on proper alignment as well as algorithm learning periods. | # '''Verification of the data''' preprocessing: Accurate alignment between animal identifiers and sensor data is critical. Errors such as duplicate device assignments to one animal (or vice versa), broken sensors, and time zone mismatches must be identified and corrected, if possible. It is recommended to consult with digital technology companies for information on proper alignment as well as algorithm learning periods. | ||
# '''Understanding the data''': This step involves identifying the type of data (e.g., raw sensor data or processed data retrieved from interfaces), its nature including units and whether it is a single shot measurement or an aggregated value, and sampling rates. Proper data visualization is recommended to uncover patterns, distributions, or anomalies. | # '''Understanding the data''': This step involves identifying the type of data (e.g., raw sensor data or processed data retrieved from interfaces), its nature including units and whether it is a single shot measurement or an aggregated value, and sampling rates. Proper data visualization is recommended to uncover patterns, distributions, or anomalies. | ||
| Line 89: | Line 89: | ||
More details can be found in Schodl et al. (2024)<ref name="Schodl2024"/> | More details can be found in Schodl et al. (2024)<ref name="Schodl2024"/> | ||
== Part 4: Use of sensor data (focus on time series data) for genetic improvement == | |||
= Part 4: Use of sensor data (focus on time series data) for genetic improvement = | |||
=== Structure of guidelines related to rumination sensor and use in genetics === | === Structure of guidelines related to rumination sensor and use in genetics === | ||
These guidelines are intended to provide stakeholders using data from sensors in dairy farms with recommendations for recording, processing, integration, and standardization across sensors; deriving novel traits for management and breeding purposes; and genetically evaluating derived functional traits. By adhering to these recommendations, stakeholders can ensure consistent and reliable data collection, leading to improved management and breeding decisions. This specific guideline focuses on rumination sensors, which monitor cows' chewing activity to assess their health and productivity, and it is part of a series of guidelines related to the use of sensor data for dairy cattle management and breeding purposes. | These guidelines are intended to provide stakeholders using data from sensors in dairy farms with recommendations for recording, processing, integration, and standardization across sensors; deriving novel traits for management and breeding purposes; and genetically evaluating derived functional traits. By adhering to these recommendations, stakeholders can ensure consistent and reliable data collection, leading to improved management and breeding decisions. This specific guideline focuses on rumination sensors, which monitor cows' chewing activity to assess their health and productivity, and it is part of a series of guidelines related to the use of sensor data for dairy cattle management and breeding purposes. | ||
For genetic purposes, rumination time has been evaluated as a proxy of feed efficiency (Byskov et al., 2017; Martin et al., 2021) and functional traits such as metabolic diseases and claw health (Moretti et al., 2017). However, there is limited information in the literature highlighting the value of rumination time as an auxiliary trait. In addition to average rumination time over specific periods, there is a growing interest in using longitudinal measurements of rumination time to define overall resilience (defined as the ability of an animal to be minimally affected by environmental disturbances and also to quickly return to its normal state). | For genetic purposes, rumination time has been evaluated as a proxy of feed efficiency (Byskov et al., 2017<ref>Martin, M. J., Dórea, J. R. R., Borchers, M. R., Wallace, R. L., Bertics, S. J., DeNise, S. K., Weigel, K. A., & White, H. M. (2021). Comparison of methods to predict feed intake and residual feed intake using behavioral and metabolite data in addition to classical performance variables. Journal of Dairy Science, 104(8), 8765–8782. https://doi.org/https://doi.org/10.3168/jds.2020-20051</ref>; Martin et al., 2021<ref>Martin, M. J., Dórea, J. R. R., Borchers, M. R., Wallace, R. L., Bertics, S. J., DeNise, S. K., Weigel, K. A., & White, H. M. (2021). Comparison of methods to predict feed intake and residual feed intake using behavioral and metabolite data in addition to classical performance variables. Journal of Dairy Science, 104(8), 8765–8782. <nowiki>https://doi.org/https://doi.org/10.3168/jds.2020-20051</nowiki></ref>) and functional traits such as metabolic diseases and claw health (Moretti et al., 2017<ref>Moretti, R., Biffani, S., Tiezzi, F., Maltecca, C., Chessa, S. and Bozzi, R., 2017. Rumination time as a potential predictor of common diseases in high-productive Holstein dairy cows. Journal of Dairy Research, 84(4), 385-390.</ref>). However, there is limited information in the literature highlighting the value of rumination time as an auxiliary trait. In addition to average rumination time over specific periods, there is a growing interest in using longitudinal measurements of rumination time to define overall resilience (defined as the ability of an animal to be minimally affected by environmental disturbances and also to quickly return to its normal state). | ||
Therefore, although we recognize the potential limitations of rumination variables for direct genetic evaluations, standardizing recording and data editing could facilitate the comparison of future research results (e.g., identification of novel traits for breeding purposes). Furthermore, rumination variables might be more useful for breeding and management purposes when combined with other datasets such as sensor-based activity measures (e.g., lying, standing, eating, drinking). It has to be stated, that sensor derived traits are proxies and are not comparable with veterinarian diagnoses. | Therefore, although we recognize the potential limitations of rumination variables for direct genetic evaluations, standardizing recording and data editing could facilitate the comparison of future research results (e.g., identification of novel traits for breeding purposes). Furthermore, rumination variables might be more useful for breeding and management purposes when combined with other datasets such as sensor-based activity measures (e.g., lying, standing, eating, drinking). It has to be stated, that sensor derived traits are proxies and are not comparable with veterinarian diagnoses. | ||
| Line 194: | Line 191: | ||
The main trait to be evaluated is “Rumination Time (min/day)”. One could consider the mean, SD, or changes in specific time windows. In addition, a new set of variables under evaluation are indicators of overall resilience based on variability in longitudinal traits, in which rumination data could be an option. Examples of these traits are rumination amplitude, log-transformed variance, and changes in rumination over time. One could also evaluate rumination patterns across lactations. Some examples of studies defining resilience based on longitudinal data are: | The main trait to be evaluated is “Rumination Time (min/day)”. One could consider the mean, SD, or changes in specific time windows. In addition, a new set of variables under evaluation are indicators of overall resilience based on variability in longitudinal traits, in which rumination data could be an option. Examples of these traits are rumination amplitude, log-transformed variance, and changes in rumination over time. One could also evaluate rumination patterns across lactations. Some examples of studies defining resilience based on longitudinal data are: | ||
* Poppe ''et al.'' (2022) | * Poppe ''et al.'' (2022)<ref name="Poppe2022">Poppe, M., H.A. Mulder, M.L. van Pelt, E. Mullaart, H. Hogeveen, and R.F. Veerkamp. 2022. Development of resilience indicator traits based on daily step count data for dairy cattle breeding. Genetics Selection Evolution 54:21. doi:10.1186/s12711-022-00713-x</ref> | ||
* Chen ''et al.'' (2023): <nowiki>https://doi.org/10.3168/jds.2022-22754</nowiki> | * Chen ''et al.'' (2023): <nowiki>https://doi.org/10.3168/jds.2022-22754</nowiki> | ||
* Poppe ''et al.'' (2021): <nowiki>https://doi.org/10.3168/jds.2020-19245</nowiki> | * Poppe ''et al.'' (2021): <nowiki>https://doi.org/10.3168/jds.2020-19245</nowiki> | ||
| Line 230: | Line 227: | ||
* To combine in an index with traditional functional traits | * To combine in an index with traditional functional traits | ||
Genetic parameters of rumination traits are presented in Brito et al. (2025): Page 10458 (ttps://doi.org/10.3168/jds.2025-26554). | Genetic parameters of rumination traits are presented in Brito et al. (2025)<ref name="Brito2025"/>: Page 10458 (ttps://doi.org/10.3168/jds.2025-26554). | ||
Open questions to follow up | Open questions to follow up | ||
| Line 242: | Line 239: | ||
* How to derive novel traits based on rumination pattern and variability? Studies are still needed. | * How to derive novel traits based on rumination pattern and variability? Studies are still needed. | ||
=== Informative references === | |||
Egger-Danner, C., I. Klaas, L. Brito, K. Schodl, J.M. Bewley, V. Cabrera, M.J. Haskell, M. Iwersen, B. Heringstad, K. Stock, A. Stygar, R. van der Linde, M. Hostens, N. Charfeddine, N. Gengler, and E. Vasseur. 2024. Improving animal health and welfare by using sensor data in herd management and dairy cattle breeding – a joint initiative of ICAR and IDF. Pages 56_63 in Proc 11th Eur. Conf. Precis. Livest. Farming, Bologna, Italy. Organizing Committee of the 11th European Conference on Precision Livestock Farming (ECPLF), University of Veterinary Medicine, Vienna, Austria | |||
Egger-Danner, C., I. Klaas, L. Brito, K. Schodl, J.M. Bewley, V. Cabrera, M.J. Haskell, M. Iwersen, B. Heringstad, K. Stock, A. Stygar, R. van der Linde, M. Hostens, N. Charfeddine, | |||
N. Gengler, and E. Vasseur. 2024. Improving animal health and welfare by using sensor data in herd management and dairy cattle breeding – a joint initiative of ICAR and IDF. Pages 56_63 in Proc 11th Eur. Conf. Precis. Livest. Farming, Bologna, Italy. Organizing Committee of the 11th European Conference on Precision Livestock Farming (ECPLF), University of Veterinary Medicine, Vienna, Austria | |||
Hogeveeen, H., Klaas, I.C., Dalen, G., Honig, H., Zecconi, A., Kelton, D.F. and Mainar, M.S. 2021. Novel ways to use sensor data to improve mastitis management. Journal of Dairy Science 104, 11317-11332. | Hogeveeen, H., Klaas, I.C., Dalen, G., Honig, H., Zecconi, A., Kelton, D.F. and Mainar, M.S. 2021. Novel ways to use sensor data to improve mastitis management. Journal of Dairy Science 104, 11317-11332. | ||
Lopes, L.S.F., Schenkel, F.S., Houlahan, K., Rochus, C.M., Oliveira Jr, G.A., Oliveira, H.R., Miglior, F., Alcantara, L.M., Tulpan, D. and Baes, C.F., 2024. Estimates of genetic parameters for rumination time, feed efficiency, and methane production traits in first lactation Holstein cows. Journal of Dairy Science, 107, 7, 4704-4713. | Lopes, L.S.F., Schenkel, F.S., Houlahan, K., Rochus, C.M., Oliveira Jr, G.A., Oliveira, H.R., Miglior, F., Alcantara, L.M., Tulpan, D. and Baes, C.F., 2024. Estimates of genetic parameters for rumination time, feed efficiency, and methane production traits in first lactation Holstein cows. Journal of Dairy Science, 107, 7, 4704-4713. | ||
=== Authors and contributors to guideline === | === Authors and contributors to guideline === | ||
These guidelines have been elaborated by the joint ICAR IDF Initiative on “Improving animal health and wellbeing by using sensor data in herd management and dairy cattle breeding” in collaboration of members of the ICAR Working Group on Functional Traits, the IDF Standing Committee of Animal Health and Welfare, international scientists, manufacturer and representatives of other ICAR bodies and stakeholders. | These guidelines have been elaborated by the joint ICAR IDF Initiative on “Improving animal health and wellbeing by using sensor data in herd management and dairy cattle breeding” in collaboration of members of the ICAR Working Group on Functional Traits, the IDF Standing Committee of Animal Health and Welfare, international scientists, manufacturer and representatives of other ICAR bodies and stakeholders. | ||
C. Egger-Danner<sup>1</sup>, I. Klaas<sup>2</sup>, L. F. Brito<sup>3</sup>, J. M. Bewley<sup>4</sup>, V. E. Cabrera<sup>5</sup>, S. Dagan<sup>6</sup>, R.H. Fourdraine<sup>7</sup>, N. Gengler<sup>8</sup>, M. Haskell<sup>9</sup>, B. Heringstad<sup>10</sup>, J. Heslin<sup>6</sup>, M. Hostens<sup>11</sup>, M. Iwersen<sup>12</sup>, F. Karlsson<sup>2</sup>, G. Katz<sup>13</sup>, M. Moleman<sup>14</sup>, M. Phelan<sup>6</sup>, E. Rossi<sup>6</sup>, K. Schodl<sup>l</sup>, D. Sieben<sup>15</sup>, K. F. Stock<sup>16</sup>, A. Stygar<sup>17</sup>, E. Vasseur<sup>18</sup>, Manufacturer representatives | C. Egger-Danner<sup>1</sup>, I. Klaas<sup>2</sup>, L. F. Brito<sup>3</sup>, J. M. Bewley<sup>4</sup>, V. E. Cabrera<sup>5</sup>, S. Dagan<sup>6</sup>, R.H. Fourdraine<sup>7</sup>, N. Gengler<sup>8</sup>, M. Haskell<sup>9</sup>, B. Heringstad<sup>10</sup>, J. Heslin<sup>6</sup>, M. Hostens<sup>11</sup>, M. Iwersen<sup>12</sup>, F. Karlsson<sup>2</sup>, G. Katz<sup>13</sup>, M. Moleman<sup>14</sup>, M. Phelan<sup>6</sup>, E. Rossi<sup>6</sup>, K. Schodl<sup>l</sup>, D. Sieben<sup>15</sup>, K. F. Stock<sup>16</sup>, A. Stygar<sup>17</sup>, E. Vasseur<sup>18</sup>, Manufacturer representatives | ||
* ''<sup>1</sup> ZuchtData EDV-Dienstleistungen GmbH, Dresdner Str. 89, 1200 Vienna, Austria,'' | * ''<sup>1</sup> ZuchtData EDV-Dienstleistungen GmbH, Dresdner Str. 89, 1200 Vienna, Austria,'' | ||
* ''<sup>2</sup> DeLaval International AB, Gustaf de Lavals Väg 15, 14721 Tumba, Sweden,'' | * ''<sup>2</sup> DeLaval International AB, Gustaf de Lavals Väg 15, 14721 Tumba, Sweden,'' | ||
| Line 305: | Line 267: | ||
* ''<sup>17</sup> Bioeconomy and Environment, Natural Resources Institute Finland (Luke), Latokartanonkaari 9, 00790 Helsinki, Finland,'' | * ''<sup>17</sup> Bioeconomy and Environment, Natural Resources Institute Finland (Luke), Latokartanonkaari 9, 00790 Helsinki, Finland,'' | ||
* ''<sup>18</sup> McGill University, Ste Anne de Bellevue, H9X 3V9, QC Canada.'' | * ''<sup>18</sup> McGill University, Ste Anne de Bellevue, H9X 3V9, QC Canada.'' | ||
[[File:Section . Figure 3.jpg|center|frame|'''Organisations of the Authors of the Guidelines Section 7.7''']] | |||
----[CE1]To be added | ----[CE1]To be added | ||
Latest revision as of 09:38, 12 March 2026
Sensor based behavior information for functional traits with focus on rumination
Part 1: General introduction
Background and aim of the guideline
Recent advancements in sensor technologies have significantly enhanced the capacity to technically support farmers and their advisors in monitoring the health, performance, and welfare of dairy cattle. As presented in the systematic review by Stygar et al. (2021)[1] and in other focused reviews (e.g., Hogeveen et al., 2021), a wide range of commercially available sensor systems exists and promises significant gains in the understanding and improvement of welfare in livestock. The technologies cover the spectrum from wearable devices with multiple functions (e.g., tracking of physiological parameters) to environmental sensors that monitor housing and climatic conditions, and collectively aim to provide actionable insights about animal health, reproductive status and welfare. Most wearable sensors rely on 3D accelerometers, which measure acceleration or motion to quantify cow behaviour. Sensor technology providers use algorithms and pattern recognition to enhance the raw accelerometer data and produce sensor systems which recognize rumination, eating, lying, standing, and other behaviours, using the data from sensors on the cow’s leg, neck, ear, or tail. The integration of sensor systems in livestock farming presents numerous opportunities to enhance animal health, performance and welfare, supporting farmer decision making on individual cow and group level and farm efficiency. However, while large amounts of sensor data are being collected, only a small fraction is currently used on farms, in genetic evaluation and breeding programs, or along the dairy value chain (Brito et al., 2025)[2]. To increase confidence in the use of data from advanced technologies and sensor-based herd management systems among key stakeholders (farmers and consultants, authorities, dairy processors, breeding and genetics organizations, and consumers), sensor-derived data need to be combined with routinely recorded data. At present, only a low fraction of commercially available sensor systems is independently validated for welfare assessment (14%; Stygar et al., 2021)[1], and beyond farmers’ own experience, few studies investigated the performance of some sensor systems in diverse farming environments, across different farm and management systems and geographical locations. These challenges motivate the need for coordinated guidance on how to define, process, and use sensor-derived behavioural information.
Against this background, the International Committee of Animal Recording (ICAR) and the International Dairy Federation (IDF) started a joint initiative aiming at improved usability of data across sensor systems and applications. The initiative leaders are the ICAR Functional Traits Working Group (ICAR FTWG) and the IDF Standing Committee of Animal Health and Welfare (IDF SCAHW) in collaboration with international experts from academia and industry organizations. The primary aim of this initiative is to promote the integrated use of sensor data and derived novel traits along the dairy value chain. Standardisation and harmonisation will be supported through guidelines that include basic definitions and recommendations regarding data processing and use. Priorities of work are based on results from a survey with manufacturers and feedback on stakeholder needs. These are:
- Establishing a common agreement on definitions and terminology for health conditions and behaviours measured with sensor systems
- Developing standards and recommendations to facilitate exchange of data and information across different farms and sensor technologies in accordance and collaboration with other ICAR standards and working groups
- Make guidelines based on best practices for data collection, handling and analysis for different use, e.g. genetics, health and welfare monitoring
- Generating recommendations, guidance and protocols for testing and self calibrating the performance of sensor systems for voluntary use
The work was started with focus on the sensor based behavioural trait rumination, which serves as a starting point; the same framework is intended to be extended to other sensor-based behavioural traits in future iterations of the guideline.
7.1.2.1 Description of data and data sources
The current guideline has the focus on data from sensor systems measuring animal behaviours. These sensor systems can provide information on behavioural measurements like rumination, eating, lying or indexes like activity indexes or alarms like for calving, in heat or health events. Various sensor systems are based on different technologies using different algorithms and provide different information to the farmer.
Part 2: Terminology
Suggested Key Performance Indicators (KPIs) for sensor-based rumination data
• Total daily rumination time in minutes per day
• Proportion of time spent ruminating per day
• Rumination time per time unit to enable investigation of circadian patterns and deviance, e.g. daily, hourly or 2-hourly summaries
• Coefficient of variation of hourly rumination
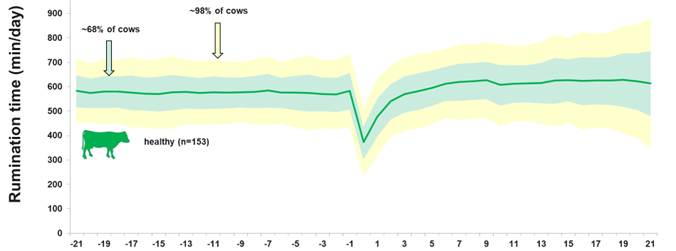
Same principle does apply to other behavioral traits like eating,..
Informative Readings
Nørgaard, P. (2003) OPtagelse af foder og drovtugning. in: Kvægets ernæring og fysiologi
Bind 1 - Næringsstofomsætning og fodervurdering. DJF rapport. Editors: T. Hvelplund and P. Nørgaard
Ruckebusch, Y. 1988. Motility of the gastro-intestinal tract. Pages 64–107 in The Ruminant Animal: Digestive Physiology and Nutrition. D. C. Church, ed. Prentice-Hall, Englewood Cliffs, NJ.
Rutter, M., (2000). Graze: A program to analyse recordings of the jaw movements of ruminants. Behavior Research Methods, Instruments and Computers 32 (1), 86-92.
Schirmann, K., von Keyserlingk, M.A.G., Weary, D.M., Veira, D.M., and Heuwieser, W (2009). Technical note: Validation of a system for monitoring rumination in dairy cows. J. Dairy Sci. 92 :6052–6055. doi: 10.3168/jds.2009-2361
Welch, J. G. 1982. Rumination, particle size and passage from the rumen. J. Anim. Sci. 54:885–894. https:// doi .org/ 10 .2527/ jas1982.544885x.
Part 3: Sensor data cleaning
Recommendations for data cleaning
These recommendations are general guidelines for understanding the data, regardless of the quality management measures implemented by the sensor technology provider. A similar approach is also used for other data e.g. in genetic evaluation.
Summary - steps for data cleaning
• Optional Sensor ICAR Device reference ID
• Validate the data merging process
• Get to know your data
• Check completeness of data
• Evaluate plausibility of sensor measures
• Detect and remove outliers
• Check for technology related noise
• Document your approach
• Outline context and purpose of further use of data
The items in this summary checklist correspond to and summarise the five-step framework described below and are intended as a quick user guide to the more detailed explanations.
Five-step framework for cleaning sensor data including
These instructions are proposed by Schodl et al. 2024[3].)
- Verification of the data preprocessing: Accurate alignment between animal identifiers and sensor data is critical. Errors such as duplicate device assignments to one animal (or vice versa), broken sensors, and time zone mismatches must be identified and corrected, if possible. It is recommended to consult with digital technology companies for information on proper alignment as well as algorithm learning periods.
- Understanding the data: This step involves identifying the type of data (e.g., raw sensor data or processed data retrieved from interfaces), its nature including units and whether it is a single shot measurement or an aggregated value, and sampling rates. Proper data visualization is recommended to uncover patterns, distributions, or anomalies.
- Checking data completeness: Missing data causing gaps in time series is a common issue and often caused by sensor malfunctions, low battery life, or poor connectivity. Depending on the subsequent analyses, missing data may require interpolation, imputation, or exclusion. Conversely, duplicate or inconsistent timestamps should be resolved to maintain data integrity. The choice between interpolation, imputation, or exclusion of missing data should be guided by the intended application, with more conservative rules recommended for genetic evaluation than for descriptive herd-level monitoring.
- Evaluating data plausibility and outlier detection: This step may be the most critical one requiring well-considered decisions by the data user. Outlier detection may be based on biological meaningful ranges, including, where possible, illustrative numeric examples (for example, typical daily rumination ranges under normal conditions), cross-checks using additional information, if available, statistical thresholds (e.g., ±3 standard deviations from the mean), and advanced modeling techniques such as Dynamic Linear Models incorporating Kalman filters (e.g., Stygar et al., 2017) or utilizing the co-dependency of data quality and model robustness (e.g., Papst et al., 2022). Regarding the management of outliers, attention should be paid to avoid removal of genuine outliers that may hold critical insights.
- Addressing technology-related noise: Sensor drift, calibration issues, and software or hardware updates may introduce inconsistencies in the data. Information on updates and handling of drift and calibration issues by the sensor company may be helpful but not necessarily easily available. Indications to look for in the data are for example the introduction of new variables, different temporal resolutions, and sudden or persistent changes in scale. Where possible, farms or data managers are encouraged to keep a simple log of firmware or software changes, calibration events, and major hardware replacements to aid interpretation of any observed shifts in the sensor data over time.
In addition to these steps, broader aspects such as the purpose and context of data analyses and the thorough documentation and transparency of the process, which are largely underreported, are essential. For instance, data for applications in herd management may have different requirements than those for genetic evaluation. As an example, if different versions of a software were used in a certain farm, but all animals from the same contemporary group had the same sensor version, the data would be useful for genetic purposes as geneticists are interested in differences among animals from the same group instead of the absolute values per se. Specific information related to data cleaning for different applications are found in the description of the use cases below.
7.3.4 Specific aspects related to the example rumination:
- To check the measured trait and confirm that it is within biological ranges (e.g. if rumination values summed up to 24-h intervals are within biologically possible estimates).
- To check for outliers caused by missing observations – this step is crucial for highly aggregated values (sums of daily observations). The activity budget of an animal (e.g. rumination, eating, and other behaviors that is not rumination or eating) should sum up to close to 24 hours. If the sum of these activities is below 20 h, it can be assumed that there was a connection problem and data were not properly stored for that 24-interval. Therefore, this observation should be removed as an outlier.
- Remove all observations from the “learning period” – (14 days, adjustable if manufactured provides evidence) after deployment of the sensors or software update (based on communication with the sensor producer or information from farmer). The “Learning period” principle should also be used when switching sensors between animals. If the learning period data is already removed by the data provider, this information should be recorded, including the length of the learning period.
- To check the number of observation days for each individual animal (with unique animal ID) – for genetic evaluation purpose we recommend a minimum number of days of data collection to be defined based on the purpose of the data use (data from certain lactation stages may be more relevant – e.g., early-lactation diseases).
More details can be found in Schodl et al. (2024)[3]
Part 4: Use of sensor data (focus on time series data) for genetic improvement
These guidelines are intended to provide stakeholders using data from sensors in dairy farms with recommendations for recording, processing, integration, and standardization across sensors; deriving novel traits for management and breeding purposes; and genetically evaluating derived functional traits. By adhering to these recommendations, stakeholders can ensure consistent and reliable data collection, leading to improved management and breeding decisions. This specific guideline focuses on rumination sensors, which monitor cows' chewing activity to assess their health and productivity, and it is part of a series of guidelines related to the use of sensor data for dairy cattle management and breeding purposes.
For genetic purposes, rumination time has been evaluated as a proxy of feed efficiency (Byskov et al., 2017[4]; Martin et al., 2021[5]) and functional traits such as metabolic diseases and claw health (Moretti et al., 2017[6]). However, there is limited information in the literature highlighting the value of rumination time as an auxiliary trait. In addition to average rumination time over specific periods, there is a growing interest in using longitudinal measurements of rumination time to define overall resilience (defined as the ability of an animal to be minimally affected by environmental disturbances and also to quickly return to its normal state).
Therefore, although we recognize the potential limitations of rumination variables for direct genetic evaluations, standardizing recording and data editing could facilitate the comparison of future research results (e.g., identification of novel traits for breeding purposes). Furthermore, rumination variables might be more useful for breeding and management purposes when combined with other datasets such as sensor-based activity measures (e.g., lying, standing, eating, drinking). It has to be stated, that sensor derived traits are proxies and are not comparable with veterinarian diagnoses.
To establish recording and data collection for rumination sensor data use in genetics, the following information is needed:
7.4.2 Required information
The items listed in Sections 1–4 below are considered essential inputs for routine genetic evaluation, whereas the fields under "Other potentially relevant information" and "Optional Information" are recommended primarily for research or extended applications when available.
The next section defines the data and standards recommended to be used for genetic evaluation. Specifications for data exchange are documented in https://github.com/adewg/ICAR.
7.4.2.1. Animal Information:
- Unique Animal ID:
- Use the ICAR ADE format (several identifier formats are accepted): Breed + Country + Sex + Identification number
- Refer to ICAR guidelines
- Recommendation: Parties exchanging data will agree on the data format for a unique Animal ID.
- For genetic evaluation it is recommended to work with farms using a herd management system and where there is the link to a national ID. A cross-reference table with link from sensor ID to different IDs on the farm including the national ID might be helpful.
- Requirements to participating farms: farmer must make sure that there is link from the sensor to a unique animal ID
- Although not recommended, sensors (and 15-digit RFID-tags) might be reused on different animals over this cannot be avoided, we recommend farmers to record this information for further verification.
- Breed:
- Use ICAR/Interbull breed codes
- Refer to breed codes
- Recommendation: Parties exchanging data need to agree on the breed codes to be used
- Lactation Number (available from other sources, e.g. DHI)
- Calving Date (from other sources):
- Format as YYYY-MM-DD
7.4.2.2. Farm Information:
- Farm ID and Site ID (use ICAR ADE standards)
- Location:
- Postal code, city, state/province, country, time zone
7.4.2.3. Sensor Information
- Sensor brand:
- Sensor type: (e.g., based on accelerometers, acoustics)
- Sensor version (or update): 20/11/2024 – not possible
- Recommendation: Data quality assurance is important for modeling in genetic evaluations. If major changes and updates were implemented in the software or sensors (and the same updates did not happen for all sensors within a farm), it is important to report this information to facilitate interpretation of the data and improve the accuracy of the genetic evaluations.
- Sensor Unique ID (not required as linked to animal ID)
- Comment: If the same sensor would be used on different animals, it is important that the information provided enables detection. Due to link to unique animal ID it should be okay and not needed. We need to be aware that problems could happen.
- Sensor ICAR Device reference ID: 8 digit identifier
- It is part of other efforts within ICAR where manufacturers can obtain an ID for some type of device they are offering to customers.
7.4.2.4. Rumination Data
- Rumination Time:
- Common basic agreement: aggregated summary of total minutes per animal per day for routine data exchange. If data of higher granularity are needed for specific purposes, such exchanges require specific agreements between the parties involved.
- Unit: min/day
- Date/Timestamp: YYYY-MM-DD (for aggregated daily values, we suggest indicating the time period summarized for example, from 00:00 to 24:00 h)
- Total daily number of minutes with measurements for rumination: When providing daily summaries of rumination per individual cow, the receiver of the data will need more information about the data editing and handling of missing values and the completeness of the shared data. Therefore, to ensure data reliability and enable broader applications, completeness indicators (e.g., number of data points collected per day, duration of each session) should also be provided. This applies also to other animal based behavior sensor information.
- Data of higher granularity (e.g. aggregated values in minutes per hour (min/h), minutes per 2 hours – min/2h) would be needed for estimating the effect of circadian patterns. Such data exchange may require specific agreements between parties for specific projects.
Data sharing for other activity parameters which can be measured in minutes
All agreed that the required information listed above for rumination applies to other behavioural traits that are measured in minutes, e.g., which can be measured in minutes like eating, lying as well as the other arrangements for rumination like total daily number of measurements.
Other potentially relevant information for genetic evaluations include the following points
Index information and alarms
- Animal-ID
- Alarm date
- Description or name of the index, which should specify how much information it represents and its main purpose, such as heat detection, calving, health monitoring, or feeding behavior assessment. It should also indicate the source of information, for example, whether it is derived from activity data, drinking behavior, or other sensor-based measures. In addition, the resolution or frequency of data collection should be described, such as whether the index is calculated on a daily, hourly, weekly, or event-based basis. Scale or coding (e.g., +/++/+++; 0/1/2; percentage; probability; mean/std dev; standardized values).
Note: there are nearly no studies using alarms for genetic analyses.
Optional Information
- Data from rumination based or related sensors:
- Frequently-collected eating time and activity level (required for some purposes – see data cleaning section)
- Alarms (e.g., heat detection, calving, disease) and indexes (health, activity, …) (see above)
- It is also worth emphasizing that other data sources will be needed (or very valuable) for genetic evaluations, including reproduction data (e.g., heat and insemination dates), health events, information on housing, milking system, grazing, feeding group, and milk yield traits (daily or per milking event).
Additional information at sensor brand level of interest
The following aspects should be documented and clarified for each sensor brand or system used:
- Animal identification: Indicate whether the animal ID can be populated using the official DHIA or herdbook identification number, or if a native link to these identifiers can be established.
- Data aggregation: Specify the number of valid data points that are aggregated within a given period (e.g., daily values), noting that this may vary by sensor brand or model.
- Sensor placement: Describe where the sensor is attached on the animal’s body, including whether it is positioned on the left or right side, as this may influence measurements and depend on the brand or sensor type.
- Handling of missing information: Provide details on how missing information is managed when calculating aggregated rumination time or other behavioral metrics.
- Interpretation of null and zero values: Clarify the meaning of null or zero values in the dataset to ensure consistent data interpretation.
- Trait documentation: Include documentation describing the traits measured, their corresponding units, the definition of indices (e.g., rumination index), and whether reported values represent sums or averages per session. Explain how missing values are handled — whether through imputation or exclusion from further processing.
- Computation of reported values: Describe the algorithm or calculation procedure used to derive reported rumination or behavioral values, including how data from individual sessions are summarized (if available).
- User-defined thresholds: Indicate whether users or farmers can set thresholds (e.g., for alerts or alarms) and whether these user-defined settings affect the data outputs provided by the system.
Before performing genetic analyses of rumination traits, one should perform descriptive statistics of the data after data processing, including minimum, maximum, mean, and standard deviation. Rumination time is widely variable depending on various factors such as diet composition, milk production level, breed, parity, lactation stage, and production system.
For breeding purposes, the main goal is to use rumination time as an auxiliary trait for improving functional traits. Therefore, for assessing the value of rumination time for use in genetics, we need to integrate rumination time records with other datasets such as other activities, health records, calving/insemination dates, and feed intake variability.
Trait definitions
The main trait to be evaluated is “Rumination Time (min/day)”. One could consider the mean, SD, or changes in specific time windows. In addition, a new set of variables under evaluation are indicators of overall resilience based on variability in longitudinal traits, in which rumination data could be an option. Examples of these traits are rumination amplitude, log-transformed variance, and changes in rumination over time. One could also evaluate rumination patterns across lactations. Some examples of studies defining resilience based on longitudinal data are:
- Poppe et al. (2022)[7]
- Chen et al. (2023): https://doi.org/10.3168/jds.2022-22754
- Poppe et al. (2021): https://doi.org/10.3168/jds.2020-19245
Factors influencing rumination time
Various factors can influence rumination time. For instance, the production system adopted in the herd such as access to grazing and outdoors space, housing type, milking system (e.g., parlors, automated milking systems), feeding system (diet, feeding group), and how/where the device is attached to an animal. For genetic purposes, we can account for these sources of phenotypic variation by fitting these effects in the genetic models as described below. The rumination sensors should be attached to the cows prior to calving (or at least shortly after calving), especially to capture potential incidence of metabolic diseases that are more frequent in early lactation. One also needs to define a “learning period” (burn-in) after the sensors are attached to the cows.
Genetic models
The main non-genetic (fixed/systematic) effects to be included in the genetic models are: a concatenation of sensor type and version/update; housing system, milking system, and feeding system (individual effects, concatenated, or by fitting contemporary group effect); Age*Parity; calving month-year; Herd*year *season (as fixed or random depending on size of farms); DIM; and number of days open. The main random effects are: herd-measurement date (day of measurement within herd) to cover impact of farm and day; and the common random effects such as additive genetic, permanent environmental, and residual effects.
Challenges / Tricky points
- There are many different sensors (and of different versions/models) being used for recording rumination-related variables, each measuring different parameters;
- How to link rumination data with functional traits (so far, no clear genetic correlations with functional traits and not enough studies)?
- Combining data from different sensor systems in genetic evaluations present challenges:
- Additional studies are needed to assess whether traits derived from different sensors are highly genetically correlated (i.e., represent the same trait).
- Clear recommendations should be provided to genetic evaluation centers.
- If trait definitions are similar and high genetic correlations across sensors are demonstrated, rumination measures may be treated as a single trait across sensors, with sensor type and/or version included as fixed or random effects in the genetic model.
- If traits derived from different sensors are not highly genetically correlated, it may be preferable to consider sensor-specific traits (e.g., in a multi-trait model) or to combine them through a selection sub-index rather than forcing them into a single trait definition.
- Data governance (GDPR/confidentiality) as multi-country genetic data sharing requires clear legal pathways
Additional points to consider
- We need to derive traits based on data from different sensors (e.g., from different companies) and estimate their variance components and genetic parameters, including genetic correlations among themselves and with other routinely-measured traits (e.g., health, performance);
- The inclusion of rumination time in a selection index will depend on the usefulness of the trait as an auxiliary trait, which is still unclear at this time;
- There is a need for evaluating the genetic correlation of rumination time across lactations as they might have different genetic background; and,
- If heifers have rumination time data (will also happen if sensors are attached prior to calving), we suggest evaluating them as separate traits (heifer and cow traits)
Taken together, the challenges and additional points listed above define priority research topics for the next phase of work and are a key reason for keeping these guidelines as a living, evolving document that can be updated as multi-brand, multi-country data accumulate.
How to combine data from sensors with traditional recording / functional traits?
- Separate
- To combine in an index with traditional functional traits
Genetic parameters of rumination traits are presented in Brito et al. (2025)[2]: Page 10458 (ttps://doi.org/10.3168/jds.2025-26554).
Open questions to follow up
- How to consider cows culled before a minimum number of days with rumination data? We believe we should keep this data, but it may be dependent on the data analysis purpose.
- How to integrate data collected in different lactation stages? (incomplete lactations)
- How to combine data from different sensor brands? Evaluate genetic correlations based on rumination traits derived from different sensor type datasets
- Could we observe less differences across sensors than data from other sensors (e.g. activity)?
- How to standardize the data from different sensors? (e.g., standardization based on mean and variance)
- Is there a value in using records from heifers?
- How to derive novel traits based on rumination pattern and variability? Studies are still needed.
Informative references
Egger-Danner, C., I. Klaas, L. Brito, K. Schodl, J.M. Bewley, V. Cabrera, M.J. Haskell, M. Iwersen, B. Heringstad, K. Stock, A. Stygar, R. van der Linde, M. Hostens, N. Charfeddine, N. Gengler, and E. Vasseur. 2024. Improving animal health and welfare by using sensor data in herd management and dairy cattle breeding – a joint initiative of ICAR and IDF. Pages 56_63 in Proc 11th Eur. Conf. Precis. Livest. Farming, Bologna, Italy. Organizing Committee of the 11th European Conference on Precision Livestock Farming (ECPLF), University of Veterinary Medicine, Vienna, Austria
Hogeveeen, H., Klaas, I.C., Dalen, G., Honig, H., Zecconi, A., Kelton, D.F. and Mainar, M.S. 2021. Novel ways to use sensor data to improve mastitis management. Journal of Dairy Science 104, 11317-11332.
Lopes, L.S.F., Schenkel, F.S., Houlahan, K., Rochus, C.M., Oliveira Jr, G.A., Oliveira, H.R., Miglior, F., Alcantara, L.M., Tulpan, D. and Baes, C.F., 2024. Estimates of genetic parameters for rumination time, feed efficiency, and methane production traits in first lactation Holstein cows. Journal of Dairy Science, 107, 7, 4704-4713.
Authors and contributors to guideline
These guidelines have been elaborated by the joint ICAR IDF Initiative on “Improving animal health and wellbeing by using sensor data in herd management and dairy cattle breeding” in collaboration of members of the ICAR Working Group on Functional Traits, the IDF Standing Committee of Animal Health and Welfare, international scientists, manufacturer and representatives of other ICAR bodies and stakeholders.
C. Egger-Danner1, I. Klaas2, L. F. Brito3, J. M. Bewley4, V. E. Cabrera5, S. Dagan6, R.H. Fourdraine7, N. Gengler8, M. Haskell9, B. Heringstad10, J. Heslin6, M. Hostens11, M. Iwersen12, F. Karlsson2, G. Katz13, M. Moleman14, M. Phelan6, E. Rossi6, K. Schodll, D. Sieben15, K. F. Stock16, A. Stygar17, E. Vasseur18, Manufacturer representatives
- 1 ZuchtData EDV-Dienstleistungen GmbH, Dresdner Str. 89, 1200 Vienna, Austria,
- 2 DeLaval International AB, Gustaf de Lavals Väg 15, 14721 Tumba, Sweden,
- 3 Department of Animal Sciences, Purdue University, West Lafayette, IN, 47907, USA,
- 4 Holstein Association USA, 1 Holstein Place, PO Box 808, VT 05302-0808 Brattleboro, United States,
- 5 University Wisconsin-Madison, 1675 Observatory Dr., WI53706 Madison, United States,
- 6 Allflex Europe sas (Allflex Europe SAS), Zl De Plague, 35510 Vitre, France,
- 7 Dairy Records Management Systems, NC State University, 313 Chapanoke Road, Suite 100, Raleigh NC 27603, USA
- 8 TERRA Teaching and Research Center, Gembloux Agro-Bio Tech, University of Liège, 5030 Gembloux, Belgium,
- 9 SRUC (Scotland’s Rural College), West Mains Road, Edinburgh EH9 3JG, United Kingdom,
- 10 Norwegian University of Life Sciences, Ås, Norway,
- 11 College of Agriculture and Life Sciences, Cornell University, 272 Morrison Hall, Ithaca, New York
- 12 Centre for Veterinary Systems Transformation and Sustainability, Clinical Department for Farm Animals and Food System Science, University of Veterinary Medicine, Veterinärplatz 1, Vienna, Austria
- 13 Afimilk LTD Afikim Israel 1514800, Israel,
- 14 Nedap Livestock, Parallelweg 2, 7141 DC Groenlo, The Netherlands,
- 15 Cowmanager B.V, Gerverscop 9, 3481 LT Harmelen, The Netherlands
- 16IT Solutions for Animal Production (vit), Heinrich-Schroeder-Weg 1, 27283 Verden, Germany,
- 17 Bioeconomy and Environment, Natural Resources Institute Finland (Luke), Latokartanonkaari 9, 00790 Helsinki, Finland,
- 18 McGill University, Ste Anne de Bellevue, H9X 3V9, QC Canada.

[CE1]To be added
- ↑ 1.0 1.1 Stygar, A.H., Gómez, Y., Berteselli, G.V., Dalla Costa, E., Canali, E., Niemi, J.K., Llonch, P., Pastell, M. 2021. A systematic review on commercially available and validated sensor technologies for welfare assessment of dairy cattle. Frontiers in Veterinary Science 8, 177
- ↑ 2.0 2.1 Poppe, M., H.A. Mulder, M.L. van Pelt, E. Mullaart, H. Hogeveen, and R.F. Veerkamp. 2022. Development of resilience indicator traits based on daily step count data for dairy cattle breeding. Genetics Selection Evolution 54:21. doi:10.1186/s12711-022-00713-x
- ↑ 3.0 3.1 Schodl, K., Stygar, A., Steininger, F., & Egger-Danner, C., 2024a. Sensor data cleaning for applications in dairy herd management and breeding. Front. Anim. Sci., 5, p.1444948. https://doi.org/10.3389/fanim.2024.1444948.
- ↑ Martin, M. J., Dórea, J. R. R., Borchers, M. R., Wallace, R. L., Bertics, S. J., DeNise, S. K., Weigel, K. A., & White, H. M. (2021). Comparison of methods to predict feed intake and residual feed intake using behavioral and metabolite data in addition to classical performance variables. Journal of Dairy Science, 104(8), 8765–8782. https://doi.org/https://doi.org/10.3168/jds.2020-20051
- ↑ Martin, M. J., Dórea, J. R. R., Borchers, M. R., Wallace, R. L., Bertics, S. J., DeNise, S. K., Weigel, K. A., & White, H. M. (2021). Comparison of methods to predict feed intake and residual feed intake using behavioral and metabolite data in addition to classical performance variables. Journal of Dairy Science, 104(8), 8765–8782. https://doi.org/https://doi.org/10.3168/jds.2020-20051
- ↑ Moretti, R., Biffani, S., Tiezzi, F., Maltecca, C., Chessa, S. and Bozzi, R., 2017. Rumination time as a potential predictor of common diseases in high-productive Holstein dairy cows. Journal of Dairy Research, 84(4), 385-390.
- ↑ Poppe, M., H.A. Mulder, M.L. van Pelt, E. Mullaart, H. Hogeveen, and R.F. Veerkamp. 2022. Development of resilience indicator traits based on daily step count data for dairy cattle breeding. Genetics Selection Evolution 54:21. doi:10.1186/s12711-022-00713-x
